Immunotherapy biomarkers and predictors of anti-checkpoint therapy success in colorectal cancer
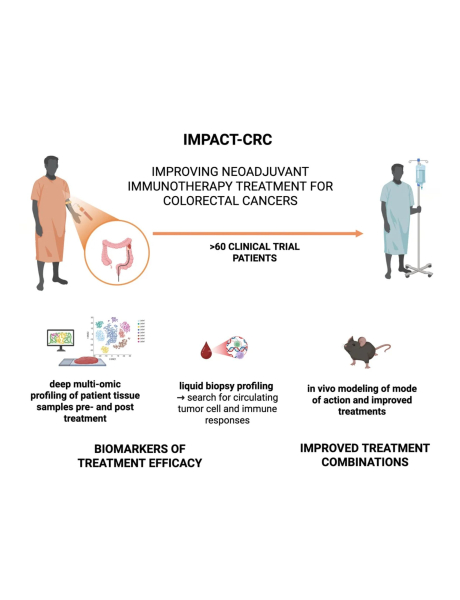
IMPACT-CRC aims to address the objectives of the EP PerMed call by leveraging neoadjuvantly treated colorectal cancer (CRC) samples and pre-clinical data to establish a robust methodology for identifying pharmacogenomic markers and signatures of immune checkpoint therapy. Using a pilot cohort of ~56 CRC samples from patients treated with the immunotherapy combination of Botensilimab (Fc-enhanced anti-CTLA4) and balstilimab (anti-PD1) (bot/bal), we will integrate multi-omics data (epigenomics, genomics, transcriptomics, and spatial transcriptomics) along with bioinformatic driven artificial intelligence-machine learning (AI/ML) pipelines to establish a robust integrative methodology to identify predictive biomarkers of treatment response.
This pilot study will serve as the foundation for the prospectively collected samples which are part of an extension of the UNICORN trial, to validate the methodologies as well as the derived biomarkers, helping to refine therapeutic strategies.
In parallel, we will incorporate mechanistic insights from pre-clinical studies using genetically engineered mouse models (GEMMs) treated with bot/bal, bridging the gap between pre-clinical and clinical research. This interdisciplinary approach, combining pre-clinical, translational research, and clinical outcomes, along with bioinformatic driven AI/ML will not only advance our understanding of the pharmacogenomic landscape in CRC but also improve patient outcomes by enabling personalized treatment strategies. Our work aligns with the EP PerMed objectives by emphasizing methodological innovation, fostering interdisciplinary collaboration, and focusing on real-world clinical impact.
Supported by Instituto de Salud Carlos III (ISCIII), Award No. AC25/00094, through “Líneas Estratégicas de Investigación en Salud” 2025 and the EP PerMed Partnership.
ROLE OF CNAG
Our Single Cell Genomics Group will contribute with advanced multi-omics profiling and integrative bioinformatics analyses, including single-cell RNA-seq, deep TCR sequencing, and immune repertoire characterization, to identify predictive biomarkers of response and resistance to immunotherapy in colorectal cancer. The group will also lead the integration of longitudinal immune and ctDNA data to model tumor–immune co-evolution during treatment.
MORE INFORMATION