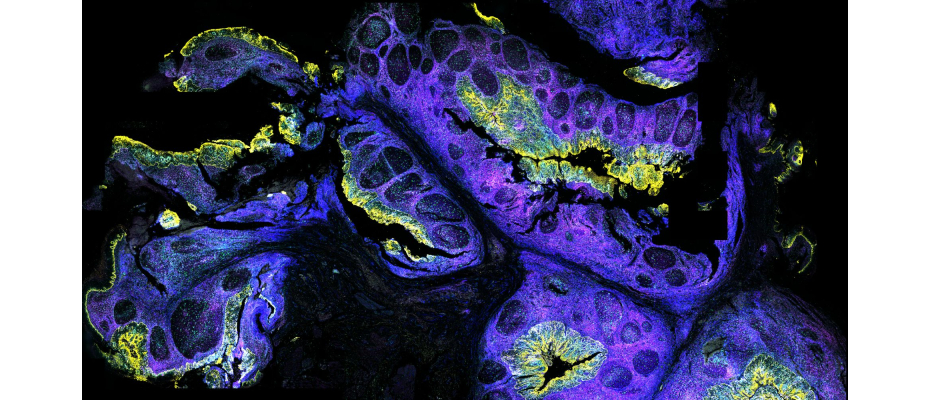
- After unveiling the cellular atlas of the human tonsil, CNAG now takes a further step with the release of its spatial transcriptomics map, advancing our understanding of this essential immune organ and revealing new defense responses in their native tissue context.
- The research, published in the European Journal of Immunology, profiles nearly 2 million cells and classifies them into 52 subpopulations across immune and structural compartments, revealing the keys to their organisation into seven distinct microanatomical niches.
- Using cutting-edge spatial genomics techniques at CNAG, researchers shed light on where biological processes occur and highlight major interaction pathways, including TNF (an inflammatory signal) and galectins (proteins that help regulate immune responses), both of which have been implicated in diverse lymphomas.
March 04, 2026. After the publication of the cellular atlas of the human tonsil (Immunity, 2024), CNAG has taken on a new challenge, building on this first approach to take a further step in understanding this key organ of our immune system. The team led by Dr Anna Pascual, our Spatial Genomics Team Leader at CNAG, with Dr Helena L. Crowell as first author, has published the spatial transcriptomics map of the human tonsil, using cutting-edge spatial genomics techniques available at CNAG, including the first CosMx instrument installed worldwide. The result, published in European Journal of Immunology and featured on its cover, is an impressive new resource that profiles nearly 2 million cells and classifies them into 52 subpopulations across immune, epithelial and stromal compartments, revealing how these diverse cell communities are arranged and work together within their microanatomical niches.
“This work shows that understanding tissues requires not only uncovering molecular and cellular composition, but also integrating spatial organisation, and cellular communication,” says Anna Pascual Reguant, senior author of the study. “Spatial genomics is opening a whole new window into our tissues, revealing layers of complexity we could only imagine before. Thanks to the spatial map of the human tonsil, we can finally reveal interactions that have remained hidden at the transcriptional level, especially those where non-immune cells play a key role”.
 More than the sum of their single-cell parts
More than the sum of their single-cell parts
Although spatial genomics offers far more than the sum of its single-cell components, detecting large numbers of RNA molecules while retaining their spatial context, its development is not yet at the same level as single-cell RNA sequencing and still lacks standardisation. For this reason, the team at CNAG developed a computational framework that integrates spatial organisation, cell–cell communication, and functional programs, establishing a versatile analytical pipeline for spatial immunology.
While previous single-cell approaches suggested that a small number of lymphoid populations dominate the tissue, spatial analysis revealed a richer diversity of epithelial, stromal, and tissue-resident immune cells, offering a more faithful representation of the organ. For example, using only scRNA-seq, the five most abundant subpopulations (largely lymphoid follicular cells) account for ~97% of all cells, whereas spatial transcriptomics shows they represent only ~70%. This approach reveals the epithelial and stromal diversity that scRNA-seq misses, as well as tissue-resident immune subsets, offering a more accurate representation of the tonsil’s cellular landscape.
Studying how cells communicate
By modelling cell–cell communication, researchers identified 181 signalling interactions across niches, including pathways such as TNF and galectin signaling, which have previously been linked to lymphoma biology. These findings highlight how mapping cells in their native environment can uncover interactions that remain hidden when cells are studied in isolation.
Together, the results provide both a comprehensive resource on tonsil organisation and a general framework for investigating how immune responses are structured within tissues. The study highlights the potential of spatial transcriptomics to uncover biological mechanisms that emerge only when cells are studied within their native context, paving the way for future research into tissue function in health and disease.
The study was carried out by the Spatial Genomics Team in collaboration with the Single Cell Genomics Group at CNAG, led by Dr Holger Heyn, and the group of Dr Elías Campo at the Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS).
SUPPORT
The research received funding from the Ministerio de Ciencia e Innovación (MCIN) under grant agreements [grant numbers PID2020-115439GB-I00, PLEC2021-007654].
REFERENCE ARTICLE
Crowell, Helena L., et al. ‘A Transcriptional Map of Human Tonsil Architecture: Beyond the Sum of (Single Cell) Parts’. European Journal of Immunology, vol. 56, no. 1, Jan. 2026, p. e70121. DOI.org (Crossref), https://doi.org/10.1002/eji.70121.











